Retrieval of Cells from Alvetex® Scaffold
● Download this protocol as a PDF (1.8 MB)
1. Introduction
The following protocol outlines a method for the retrieval of cells cultured in Alvetex Scaffold. Example data were obtained using this protocol to isolate HepG2 hepatocyte cells and A549 lung carcinoma cells cultured in Alvetex Scaffold for 7days in 6 well inserts (AVP004) in the Well Insert Holder in a Deep Petri Dish (AVP015) format.
2. Method
- Unclip inserts and carefully remove the Alvetex Scaffold discs using flat-ended forceps.
- Gently wash each disc in PBS and cut the disc into quarters with a sterile scalpel or sterile dissecting scissors to increase the exposed surface area. Transfer the quarters to a sterile 15 mL centrifuge tube containing 5 mL of 0.25 % Trypsin-EDTA (Sigma, T4049).
- Incubate the tube at 37 °C, 5 % CO2 on a shaking platform set to 100 rpm for 10 minutes.
Note: To maximise agitation, the 15 mL centrifuge tube should be held sideways on the platform rather than upright. This can be safely achieved by placing the centrifuge tube in a tube rack and then placing the rack on its side on the platform.
- Shake the tube by hand vigorously for 10 seconds to encourage cells to detach, then transfer the resulting cell suspension to a 50 mL centrifuge tube and add FBS-containing medium to neutralise the trypsin solution. Reserve the detached cells at 37 °C, 5 % CO2.
- Add a new 5 mL of 0.25 % Trypsin-EDTA to the 15 mL centrifuge tube containing the quarter discs and repeat steps 3 and 4, combining the two fractions of detached cells into the same 50 mL tube.
- Centrifuge at 1200 rpm for 5 minutes to pellet the cells.
- Resuspend the cell pellet in an appropriate volume of medium for downstream processes.
3. Example: HepG2 cells and A549 cells grown for 7 days in Alvetex Scaffold 6 well insert in Petri dish format
HepG2 cells (ATCC, HB-8065) were routinely maintained in T-75 flasks. HepG2 complete media consisted of: EMEM media (Lonza, BE12-125F) supplemented with 10 % v/v FBS, 2 mM L-glutamine and 100 U/mL Penicillin/Streptomycin. A549 cells (ATCC, CCL-185) were routinely maintained in T-75 flasks. A549 complete media consisted of: DMEM media (Lonza, BE12-614F) supplemented with 10 % v/v FBS, 2 mM L-glutamine and 100 U/ml Penicillin/Streptomycin. Cells were seeded onto Alvetex Scaffold discs in 6 well inserts (AVP004) in well insert holders in deep Petri dishes (AVP015), at a density of 1 × 106 cells per insert. Cultures were maintained for 7 days, with media changes on days 2 and 4. After 7 days, cultures were processed according to the protocol described above.
The total number and number of viable cells retrieved were counted using Trypan blue exclusion method. Alvetex Scaffold discs were analysed post-retrieval by MTT assay and histological staining to estimate cell numbers remaining in the 3D substrate.
Results
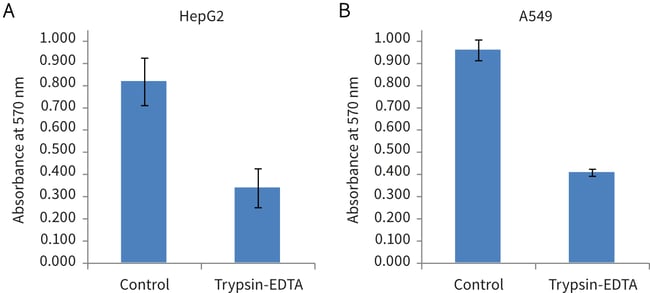
Figure 1. Biochemical analysis of cell viability using a standard MTT assay on Alvetex Scaffold discs post-cell extraction using trypsin-EDTA. Data from 4 sample replicates of HepG2 (A) and A549 (B) cultures are shown (n = 4, mean ± SEM).
On the basis of residual in-scaffold MTT assay data, 58 % of HepG2 cells and 57 % of A549 cells were retrieved (compared to untreated control discs) using the protocol described above.
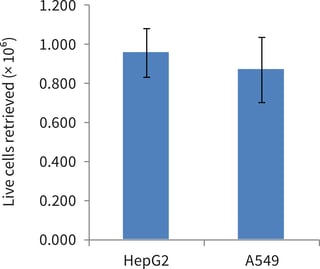
Figure 2. Trypan blue counts of live cells retrieved from single 6 well format Alvetex Scaffold discs post-cell extraction using trypsin-EDTA. Data from 3 sample replicates of HepG2 and A549 cultures are shown (n = 3, mean ± SEM).
Cell viability post-retrieval was 93 % for HepG2 cells and 96 % for A549 cells, as determined by cell counting in conjunction with Trypan blue exclusion.
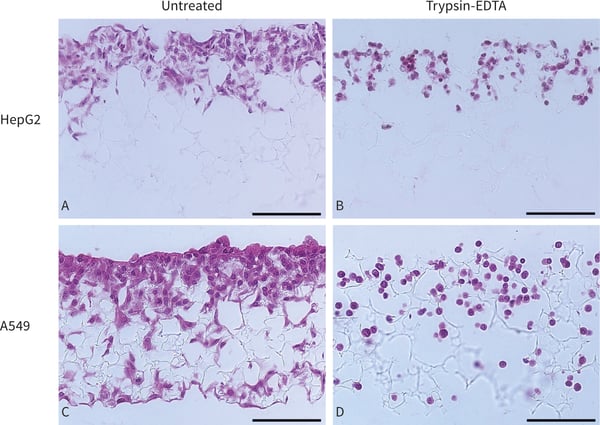
Figure 3. Bright field micrographs showing the morphology of HepG2 cells and A549 cells cultured for 7 days on Alvetex Scaffold in 6 well insert within a Petri dish format (×20 magnification, scalebars = 100 µm). (A.) untreated HepG2 cells and (B.) HepG2 cells after Trypsin-EDTA treatment. (C.) untreated A549 cells and (D.) A549 cells after Trypsin-EDTA treatment. Cells were fixed, embedded in paraffin wax, sectioned (7 µm) and counterstained with haematoxylin and eosin. The effect of trypsin action can clearly be seen on residual cells which lose contact with each other and adopt a rounded morphology. The specific images shown exhibit cell removal rates of 39 % for HepG2 cells and 50 % for A549 cells, as estimated by cell counting.