Extraction of RNA from Cells Cultured in Alvetex® Scaffold in 3D
● Download this protocol as a PDF (0.6 MB)
1. Introduction
The following protocol outlines the use of a commercial extraction kit (Qiagen RNeasy® mini kit, 74104) for the isolation of total RNA from cells cultured in Alvetex Scaffold. Example data was obtained using this protocol to extract RNA from HepG2 hepatocytes cultured in Alvetex Scaffold for 7 days in 6 well inserts (AVP004) in Well Insert Holder in Deep Petri Dish (AVP015) format.
2. Method
- Wash Alvetex Scaffold culture by gentle immersion in PBS using flat-ended forceps and transfer to a clean 12 well plate.
- Lyse cells by adding 600 µL Qiagen RNeasy® kit lysis buffer RLT per well.
- Place on a rotating platform (100 rpm) for 10 minutes at room temperature.
- Homogenise the lysate by passage up and down through a 20-gauge needle 10 times using a
sterile plastic syringe. - Add an equal volume of 70 % ethanol to the homogenised lysate and pipette up and down
10 times to mix. - Transfer sample (up to 700 µL at a time) to an RNeasy® spin column in a 2 mL collection tube. Close cap and microcentrifuge for 15 seconds at 8000 × g. Discard flow-through. Repeat for remaining volume of lysate.
Optional: On-column DNase digestion can be performed at this point (see steps 6 a-d).
Alternatively, proceed to step 7.- Add 350μL Buffer RW1 to the column. Close cap, and microcentrifuge for 15 seconds at 8000 × g to wash the membrane. Discard flow-through.
- Prepare DNase1 reagent by adding 10 ≥L DNase1 stock solution (prepared according to manufacturer’s instructions) to 70 µL Buffer RDD. Mix by gentle pipetting or gentle tube inversion.
- Add the 80 μL diluted DNase1 reagent to the column membrane and incubate at room temperature for 15 minutes.
- Add 350 µL Buffer RW1 to the column. Close cap and microcentrifuge for 15 seconds at 8000 × g. Discard flow-through. Proceed to step 8.
- Add 700 μL Buffer RW1 to the column. Close cap and microcentrifuge for 15 seconds at 8000 × g, reusing the collection tube. Discard flow-through.
- Add 500 μL Buffer RPE (appropriately diluted with ethanol according to manufacturer’s instructions) to the column. Close cap and microcentrifuge for 15 seconds at 8000 × g. Discard flow-through.
- Add a further 500 μL Buffer RPE to the column. Close cap and microcentrifuge for 2 minutes at 8000 × g. Discard flow-through.
- Transfer column to a clean 2 mL collection tube. Close cap and microcentrifuge at full speed for 1 minute.
- Transfer column to aclean 1.5mL collection tube and add 50 μL RNase-free water onto the column membrane. Close cap and leave at room temperature for 5 minutes. Microcentrifuge for 1 minute at 8000 × g to elute RNA.
3. Example: Extraction of Total RNA from HepG2 Hepatocytes Cultured in Alvetex Scaffold in 3D
3.1. Cell Culture details
HepG2 cells (ATCC, HB-8065) were routinely maintained in T-75 flasks. HepG2 complete media consisted of: MEM media (Gibco (Thermo Fisher), 21090) supplemented with 10 % v/v FBS, 2 mM L-glutamine and 100 U/mL Penicillin/ Streptomycin. Cells were seeded onto Alvetex Scaffold discs in 6 well inserts (AVP004) in Well Insert Holders in Deep Petri dishes (AVP015), at a density of 1 × 106 cells in 150 µL media suspension per insert. After settling for 3 hours in an incubator (5 % CO2, 37 °C) complete media was carefully added (70 mL per Petri dish). Cultures were maintained for 7 days, with a single media change on day 5. After 7 days, well inserts were dismantled and cultures were processed according to the protocol described above.
3.2. RNA analysis results
RNA samples were assessed for quantity and quality by spectrophotometric (Table 1) and agarose gel (Figure 1) analyses.
| Sample 1 | Sample 2 | |
|---|---|---|
| RNA yield (μg/mL) | 185.1 (± 0.044) | 182.7 (± 0.470) |
| A260/ A280 | 2.123 (± 0.012) | 2.117 (± 0.006) |
Table 1. RNA yield and purity analysis. RNA samples from duplicate HepG2 Alvetex Scaffold cultures were diluted 5-fold in 10 mM Tris-HCl, pH 7.0. Values shown represent the mean of triplicate readings per sample (± SD), taken on a NanoDrop 1000 spectrophotometer (Thermo Scientific).
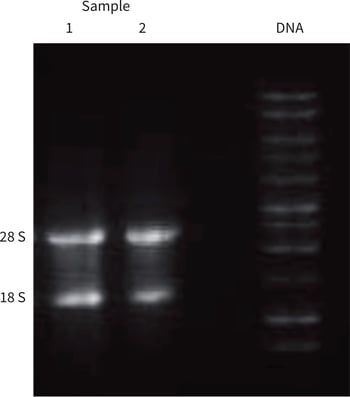
Figure 1. Agarose gel RNA profile from duplicate HepG2 Alvetex Scaffold 3D cultures (Samples 1 and 2). 28S and 18S rRNA bands are clearly visible confirming RNA integrity (1 % agarose gel; visualisation via ethidium bromide staining; 1 kb molecular weight ladder: Promega, G571A).